Diagnostic performance of electronic nose technology in chronic lung allograft dysfunction
Aim: To assess the diagnostic accuracy of exhaled breath analysis using eNose technology to detect chronic lung allograft dysfunction (CLAD) in lung transplant recipients. Additionally, the study sought to evaluate whether eNose could distinguish between different CLAD phenotypes, such as bronchiolitis obliterans syndrome (BOS) and restrictive allograft syndrome (RAS), and to examine the added diagnostic value of combining eNose data with known CLAD risk factors
Take home message: eNose technology can accurately detect chronic lung allograft dysfunction (CLAD) in lung transplant patients, distinguishing between CLAD phenotypes like BOS and RAS. This non-invasive tool shows potential as a quick, point-of-care diagnostic aid for early detection and management of CLAD.
Introduction
Chronic lung allograft dysfunction (CLAD) remains a significant challenge in lung transplantation (LTx), with limited biomarkers available for early detection. This study assessed the diagnostic value of exhaled breath analysis using the electronic nose (eNose) in identifying CLAD and differentiating its phenotypes.
Methods
A cross-sectional analysis was conducted on 152 lung transplant recipients (LTR), including 38 patients diagnosed with CLAD. Real-time exhaled breath analysis was performed using the SpiroNose® device, which utilizes multiple cross-reactive sensors to capture volatile organic compounds (VOCs) in breath samples. Data were analyzed using partial least squares discriminant analysis (PLS-DA) and logistic regression models. The study also examined the added value of combining eNose breathprints with known clinical risk factors for CLAD (e.g., age, gender, type of transplant, time post-transplant, and prior acute cellular rejection).
Results
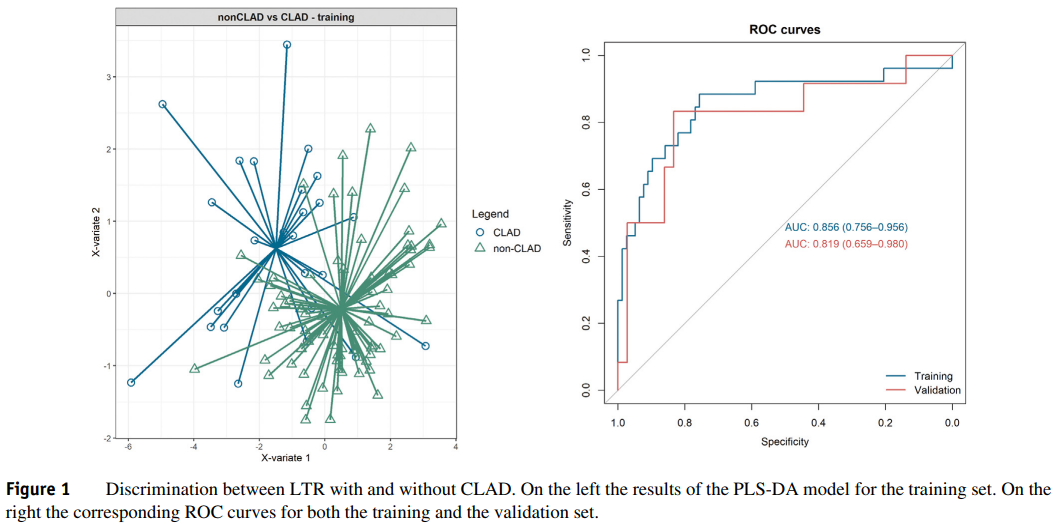
ENose breath analysis achieved a classification accuracy of 79-83% in distinguishing between LTR with and without CLAD, with an AUC of 0.86 in the training set and 0.82 in the validation set. Incorporating clinical risk factors improved the model’s accuracy, increasing the AUC to 0.94 for both training and validation sets (p < 0.05). The eNose effectively differentiated between the two main phenotypes of CLAD: bronchiolitis obliterans syndrome (BOS) and restrictive allograft syndrome (RAS), with an AUC of 0.95. However, the accuracy was lower for distinguishing other phenotypes and stages of CLAD.